Core A Software
 |
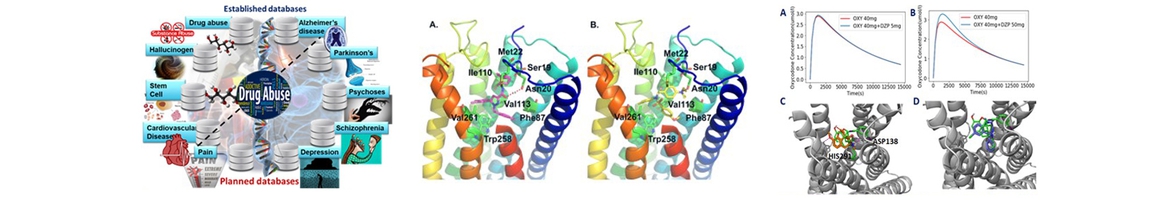
DAKB Chemogenomics Database for Drug abuse Research is designed for facilitating data-sharing and information exchange among scientific research communities for drug abuse, including genes, proteins, small molecules and signal pathways, with online structure search functions and data analysis tools implemented. http://www.cbligand.org/CDAR |
 |
TargetHunter Ligand-based target prediction tools with BioassayGeoMap function integrated. http://www.cbligand.org/TargetHunter |
 |
HTDocking Structure-based target prediction tools for drug repurpose research. http://www.cbligand.org/HTDocking/ |
 |
BBB Predictor Machine learning algorithms- AdaBoost and SVM-based tool is designed for predicting the permeability of blood-brain barrier (BBB) for compounds. http://www.cbligand.org/BBB/ |
 |
GPU-Accelerated Compound Library Comparison Modern graphics process units (GPU) –based parallel computing, millions of compound comparison can be accomplished in a few seconds. http://www.cbligand.org/gpu/ |
 |
LiCABEDS Ligand Classifier of Adaptively Boosting Ensemble Decision Stumps is designed to help pharmacologists and medicinal chemists to easily use machine learning techniques for virtual screening and quantitative structure-activity relationship (QSAR) study. http://www.cbligand.org/LiCABEDS/ |
 |
CBID A Web-interfaced cannabinoid molecular information database repository that are designed and constructed to facility data-sharing and information exchange among scientific research communities with on-line structure search functions and data analysis tools implemented. http://www.cbligand.org/CBID |
Core B Software
 |
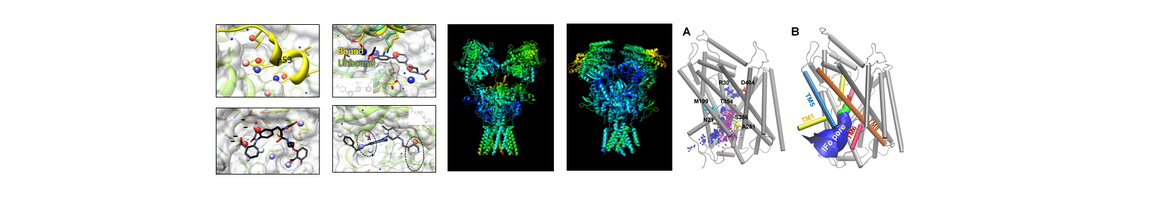
DynOmics ENM server computes biomolecular systems dynamics by integrating two widely used elastic network models (ENMs) – the Gaussian Network Model (GNM) and the Anisotropic Network Model (ANM). Unique features include the consideration of environment, the prediction of potential functional sites and reconstruction of all-atom conformers from deformed coarse-grained structures. http://enm.pitt.edu/ |
 |
iGNM 2.0 A Gaussian Network Model (GNM) Database where users may access the results for the equilibrium dynamics of any given biomolecular structure, or corresponding biological assembly, structurally characterized to date. http://ignm.ccbb.pitt.edu/ |
 |
ProDy An Application Programming Interface, for sequence- and structure-based studies of functional mechanisms and collective dynamics of biomolecular systems. It is a free and open-source Python package. It has fast, flexible and powerful file parsers and customizable atom-selections for structural analyses and for bridging sequence evolution and structural dynamics http://prody.csb.pitt.edu/ |
 |
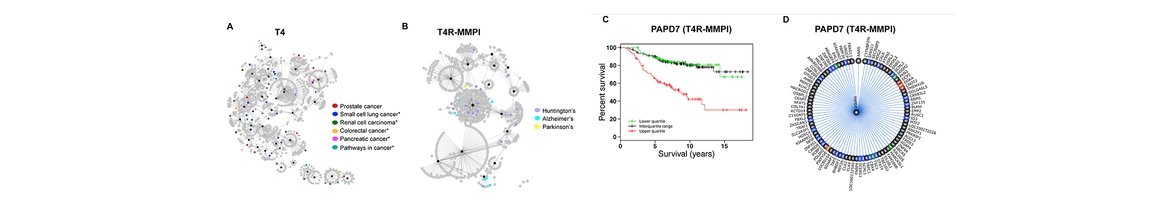
QuartataWeb Server Addresses the need for efficient identification of potential interactions between drugs and targets, and identification of repurposable drugs, or side effects of known drugs, using a latent factor model of drug-target interactions. http://quartata.csb.pitt.edu/ |
| ANM A Normal Mode Analysis (NMA) Server for analysis of collective motions of biomolecular systems using the anisotropic network model (ANM). The ANM is an elastic network model (ENM) where the system is represented by a network of nodes connected by elastic springs. It lends itself to fast and accurate evaluation of cooperative motions for large complexes and assemblies. http://anm.csb.pitt.edu/ |
|
 |
Rhapsody Rhapsody is a web tool for pathogenicity prediction of human missense variants based on sequence, structure and dynamics of proteins. http://rhapsody.csb.pitt.edu/ |
Core C Software
 |
GenAMap A visualization strategies for structured association mapping. https://github.com/blengerich/GenAMap |
 |
Precision Lasso Accounting for Correlations and Linear Dependencies in High-Dimensional Genomic Data Website |
 |
CMM Enables a joint GWAS analysis on two independently collected sets of GWAS data with different phenotypes. Website |
 |
Confounder filtering An efficient method that can remove the influences of confounding factors such as age or gender to improve the across-cohort prediction accuracy of neural networks. Website |
 |
CS-LMM Aims to uncover genetic variants of weaker associations by incorporating known associations as a prior knowledge in the model. Website |
 |
MKKC The MKKC package performs the robust multiple kernel k-means clustering using min-max optimization. This package also includes 18 multi-view simulation data generated for illustration purpose. Website |